 You can toggle the display of this translation track by clicking once, anywhere in the sequence or translation track, or by toggling Show Translation in the track popup menu. When you zoom all the way in, the amino acid symbols will appear. Methionines are colored green, and all stop codons are colored red. svg.Īmino acids are displayed as blocks colored in alternating shades of gray. Search a DNA sequence to match either a DNA query, or a protein translation, or an annotation. Selecting Save image from the right-click pop-up menu save the lower display panel containing the Sequence track as an image. Specify the image file format by setting the filename extension in the file save dialog to. Right-click on Sequence track to select Show translation from the pop-up menu and to select a Translation Table. 
A library of annotated files for common plasmids. Master SnapGene & key molecular biology & bioinformatics concepts. The translation is shown for the strand indicated. A short video series for new SnapGene users. With the reference genome sequence track, you can optionally display a 3-band track that shows a 3-frame translation of the amino acid sequence for the corresponding nucleotide sequence. This strand will show the complement nucleotides and reverse complement translations. SnapGene Viewer is a revolutionary software app that allows molecular biologists to create, browse, and share richly annotated DNA sequence files up to 1 Gbp in length. An arrow pointing left indicates that the negative strand is showing. The direction of the arrow indicates which strand is currently displayed. Note that the sequence and the arrow are only displayed when zoomed in to a sufficiently small region.Īlternatively, right-click on Sequence track to select Flip strand from the pop-up menu. You can change the strand that is displayed by clicking on the arrow in the title to the left of the track. A useful discussion about this can found on Bioinformatics StackExchange (see for example response #11 on this thread). However, the convention for the use of case and N, is not completely standardised, and depends on the creator of the genome sequence. Lower case letters often mark repeated regions, and N/n may represent ambigous nucleotides. In this tutorial, you will learn how SnapGene can help you view and edit translated features, open reading frames, and whole sequence translations. In addtion to the upper case letters A, C, G, and T, you may see lower case letters for these bases, and also N / n. IGV displays the sequence of bases as they appear in the FASTA file for the reference genome. Free software that allows you to create, browse, and share richly annotated sequence files. To change this default nucleotide coloring scheme see the Modify the prefs.properties file page. The sequence is represented by colored bars or colored letters, depending on zoom level, with adenine in green, cytosine in blue, guanine in yellow, and thymine in red ( A, C, G, T). 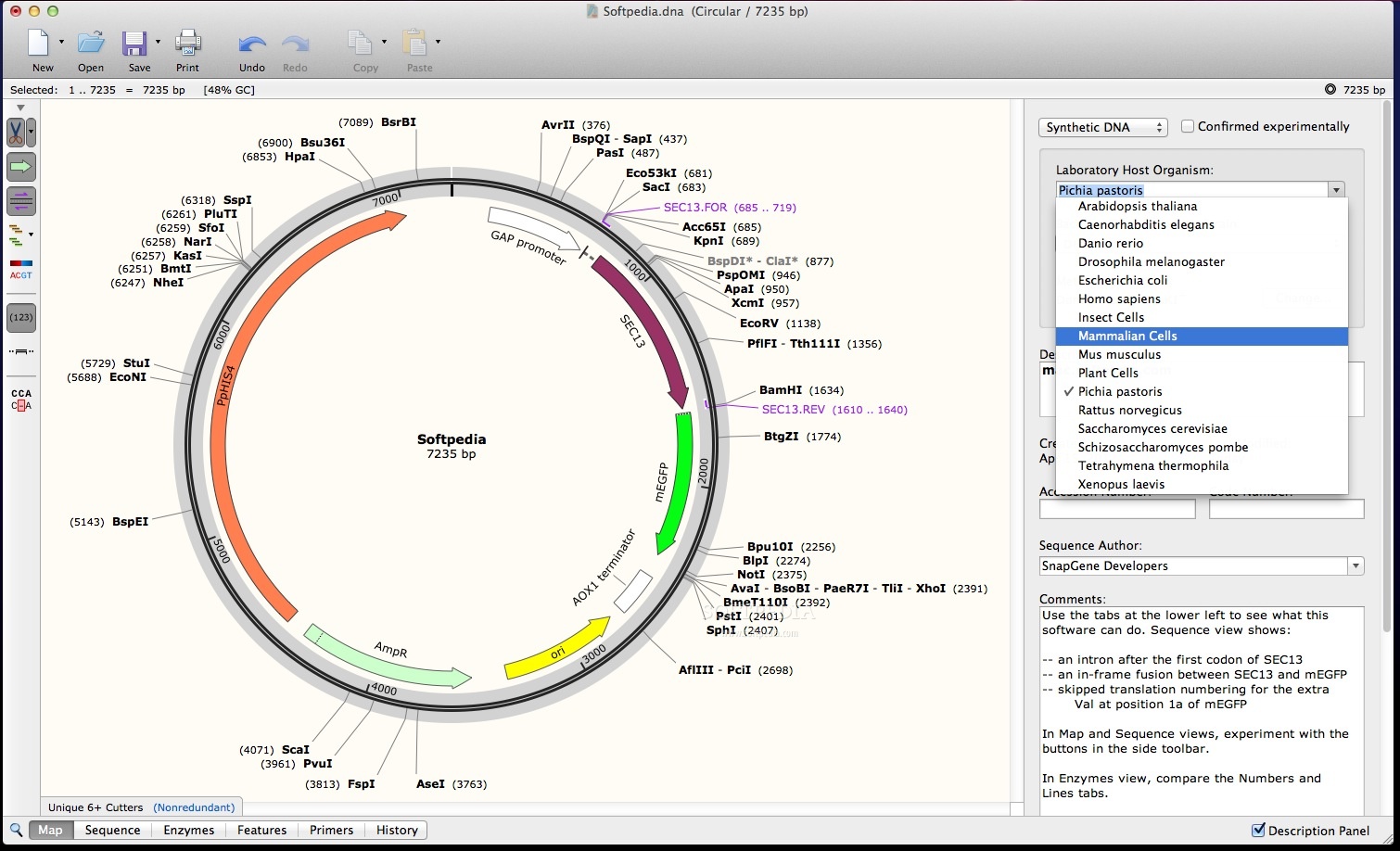
As a result, your scientists can switch entirely to SnapGene without losing data, or can continue using legacy software together with SnapGene without conflict.Īs a service to the research community, SnapGene provides tutorial videos along with a library of carefully annotated plasmids, along with guides to popular cloning methods.When zoomed in sufficiently, the reference genome Sequence track appears at the top of the lower panel above the Genes track, if any, in the IGV display as shown in the Screenshot (2015.04.01). coding sequence (you can add / remove restriction sites to your coding sequence without changing the translation). SnapGene supports a host of file formats. 
0 Comments
Leave a Reply. |
 RSS Feed
RSS Feed
